Antibiotic resistance is one of the greatest threats to public health. Each year, resistant bacterial infections kill hundreds of thousands of people worldwide, and the numbers are rising. But where does this resistance come from, and how does it spread so quickly between bacteria, even across diverse environments? The article “Genomic Analysis of IncH Plasmids Reveals Their Role as Drivers of Antimicrobial Resistance and Adaptive Traits in Enterobacterales” in FEMS Microbiology Letters identifies IncH plasmids as a potential answer, as outlined by Bradd Mendoza-Guido in this #FEMSmicroBlog. #MicrobiologyIsEverywhere
Plasmids spread antibiotic resistance between different settings
Many bacteria carry plasmids of different lengths, like portable USB drives of different sizes. These carry genes that confer survival advantages, such as antibiotic resistance.
Antibiotics widely used in clinical practice include aminoglycosides, β-lactams, and sulphonamides. These inhibit essential bacterial growth mechanisms, making these antibiotic classes effective for many bacterial infections.
Since bacteria can exchange their plasmids, their genes can spread across different species and habitats. Hence, antibiotic resistance genes found in a farm animal in China today could, in principle, reach a hospital patient in Costa Rica at some point.
The study “Genomic Analysis of IncH Plasmids Reveals Their Role as Drivers of Antimicrobial Resistance and Adaptive Traits in Enterobacterales” in FEMS Microbiology Letters focuses on a particularly concerning group called IncH plasmids.
These tend to be larger than average plasmids, which likely formed when distinct plasmids fused inside a host bacterium conferring multiple adaptive advantages. As highly mobile elements, IncH plasmids are found in the bacterial family Enterobacterales, some of which are well-known pathogens causing serious infections in humans.
IncH plasmids cause alarming levels of resistance
The study analysed 1,308 IncH plasmid sequences collected from bacteria across 59 countries from clinical and non-clinical settings, available in one of the most comprehensive databases. Based on genomic and evolutionary approaches, the authors characterized key adaptive features and delineated the genetic networks linking human and non-human environments.
The goal was to better understand antimicrobial resistance dynamics within a One Health framework, since IncH plasmids are globally distributed and predominantly associated with human pathogens. So far, they have been reported on every inhabited continent, with the highest numbers present in China, the United States, and Japan.
These plasmids are commonly found in major pathogenic species such as Salmonella enterica, Klebsiella pneumoniae, Escherichia coli, Enterobacter hormaechei, Enterobacter cloacae, and Citrobacter freundii. Some subtypes displayed distinct host associations; a relevant pattern for future epidemiological investigations.
Notably, IncH Plasmids from clinical and non-clinical sources share similar resistance gene patterns. This strongly supports the One Health view: human, animal, and environmental health are deeply interconnected, and resistance traits circulate freely among them.
Furthermore, over 70% of analysed plasmids harboured genes conferring resistance to aminoglycosides, while roughly 60% of them harboured genes conferring resistance to three or more antibiotic groups—including β-lactams and sulfonamides. Nearly one in five plasmids carried resistance to carbapenems, antibiotics considered a last resort when all others fail.
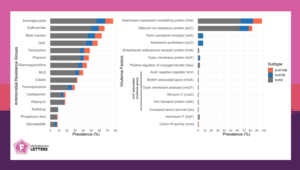
In addition to antibiotic resistance, IncH plasmids also carry genes that help bacteria survive environmental stress. Some confer resistance to metals like tellurium and may contribute to increased virulence. Hence, it seems that IncH plasmids are not just resistance vectors, but true bacterial survival enhancers.
The reason why IncH plasmids accumulate a high number of resistance genes seems to be due to the presence of integrons. Found in most IncH plasmids, they act as resistance “gene collectors” or molecular traps, while conferring resistance to carbapenems in themselves.
Plasmid and antimicrobial resistance dynamics beyond clinical settings
These findings underscore that monitoring antibiotic resistance cannot be limited to hospitals. Rivers, soils, food systems, and animal farms are all reservoirs in which resistance genes accumulate and spread. IncH plasmids, with their abilities to colonize diverse bacterial hosts across diverse environments, are key players in this global resistance crisis.
For policymakers and public health authorities, this work highlights the urgent need for integrated surveillance programs. These should monitor resistance genes (along with the plasmids and other mobile elements that carry them) both in clinical settings and the environment and food chain,adopting a true One Health approach. Understanding and tracking these molecular drivers of resistance is essential to slowing the spread of untreatable infections and protecting the antibiotics we still can use.
- Read the article “Genomic Analysis of IncH Plasmids Reveals Their Role as Drivers of Antimicrobial Resistance and Adaptive Traits in Enterobacterales” by Mendoza-Guido et al. in FEMS Microbiology Letters (2025).

Bradd Mendoza-Guido is a Ph.D. student and early-career researcher in the Laboratory of Molecular and Microbial Epidemiology of Aquatic Environments at the Health Research Institute of the University of Costa Rica. He holds a Bachelor’s degree in Biology and a Master’s degree in Bacteriology. He currently leads several projects combining bioinformatic and genomic analyses with experimental laboratory assays, aiming to understand the role of plasmids and other mobile genetic elements in bacterial adaptation, particularly their contribution to the emergence and spread of antimicrobial resistance. Bradd aims to continue demonstrating that mobile genetic elements should be regarded not as mere vectors of traits, but as molecular symbionts that interact with bacteria under the same evolutionary and ecological dynamics governing all living organisms.
About this blog section
The section #MicrobiologyIsEverywhere highlights the global relevance of microbiology. The section acknowledges that microbiology knows no borders, as well as the fact that microbiologists are everywhere and our FEMS network extends well beyond Europe. This blog entry type accepts contributions from excellent blogs translated into English. Regional stories with global relevance are welcomed. National or international events sponsored, organised or connected to FEMS are also covered.
| Do you want to be a guest contributor? |
| The #FEMSmicroBlog welcomes external bloggers, writers and SciComm enthusiasts. Get in touch if you want to share your idea for a blog entry with us! |



