Although greenhouse gases are present in the atmosphere in relatively small amounts, their increasing concentrations cause stronger heat trapping on Earth. The second most abundant greenhouse gas, methane, has a high heat-trapping capacity but a short lifespan in the atmosphere. Therefore, decreasing atmospheric methane levels would result in quicker decreases in warming. So-called “microbial methane munchers” have the unique abilities to lower methane emissions from diverse ecosystems. That’s why the article “Uncovering hidden phylo- and ecogenomic diversity of the widespread methanotrophic genus Methylobacter” in the Thematic Issue “Advances in Microbial C1 Metabolism” explores methanotrophs and methane oxidisers as an unrecognised species diversity. #FascinatingMicrobes
Methanotrophs and methane oxidisers in their diverse environments
Manipulating atmospheric methane concentrations could help us combat the disastrous effects of climate change. Methane is emitted to the atmosphere by both natural processes and human activity, with the latter increasing.
Methanotrophs or methane oxidisers present in soil, water, and sediments ‘munch on methane’: they generate energy from methane and build biomass. Through these unique metabolic abilities, they effectively lower methane emissions from different environments, for instance from wetlands and lakes.
Among methane oxidisers, the genus Methylobacter is globally widespread. In certain soils, especially in the Arctic, water columns, lake sediments, groundwater, and human-made habitats, its members often represent the most numerous methane oxidisers.
Some of these places are theoretically hostile for Methylobacter and other aerobic methane oxidisers. They are severely limited or even depleted in oxygen—a crucial resource for aerobic methane oxidation. Moreover, these places are not only extremely different from each other regarding water chemistry, available resources, temperature and more, but they are also severely affected and altered by climate change. For instance, higher temperatures and deeper permafrost thaw impact Arctic soils, while the progressive decline in oxygen concentrations affects freshwater ecosystems around the globe.
That’s why the article “Uncovering hidden phylo- and ecogenomic diversity of the widespread methanotrophic genus Methylobacter” in FEMS Microbiology Ecology aims to understand how Methylobacter can thrive and munch on methane in all these different places.
The metabolic potential of Methylobacter
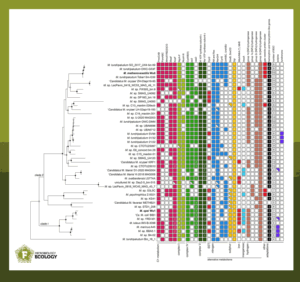
To understand their metabolic potential, the authors investigated nearly 100 genomes labelled as Methylobacter. According to the current standards of distinguishing microbial species based on their genomic DNA, the genus Methylobacter is much more diverse than previously thought.
Instead, the study found over 30 putative species; this is roughly half the number of known mammal species in the entire Czech Republic, where the research team is based. They showed that the taxonomic ‘barcode’, the sequence of the 16S rRNA gene, which reveals the different bacteria and archaea in a sample, does not provide adequate resolution to tell Methylobacter species apart.
Most importantly, the study highlights unique sets of genes in each of these species. These encode enzymes and proteins able to make use of versatile resources or the same resources in various conditions.
For instance, some of the Methylobacter species contain up to three different sets of genes encoding methane monooxygenase—the enzyme that enables methane oxidation. While the canonical and non-canonical methane monooxygenases are attached to the main system inside the cell membrane, a soluble monooxygenase not only floats in the cytoplasm but also oxidises both methane and other carbon sources. Theoretically, Methylobacter with the soluble type have access to more versatile ‘food’ sources, giving them a survival advantage in certain environments.
Some of the newly identified genes might even encode for compounds with potential use, like medicines or other useful compounds. As such, sustainable industrial processes involving greenhouse gas reduction could be coupled with microbial product formation.
These hypotheses and others need to be experimentally tested and quantified to understand whether and how these metabolic pathways impact Methylobacter’s capacity to munch on methane and lower emissions. Unfortunately, the majority of Methylobacter species have not yet been isolated and cultured, currently preventing researchers from testing these hypotheses. Therefore, modelling tools can help predict optimal growth conditions for uncultured Methylobacter species, which in turn could help researchers characterise them.
In summary, this study opens up avenues for future studies on the diverse Methylobacter species with the goal of better understand how these important methane-munchers react to climate change.
- Read the article “Uncovering hidden phylo- and ecogenomic diversity of the widespread methanotrophic genus Methylobacter” in the Thematic Issue “Advances in Microbial C1 Metabolism” by Wutkowska et al. in FEMS Microbiology Ecology (2025).

Magdalena Wutkowska is a postdoc in the Microbes CaN Cycle lab at the Biology Centre of the Czech Academy of Sciences (České Budějovice, Czechia), mentored by Dr. Anne Daebeler. In her work, shea aims to disentangle factors that modulate interactions between methane oxidisers and nitrifiers — two groups of organisms with a powerful impact on the global biogeochemistry. She also studies survival strategies and alternative metabolisms of aerobic methane oxidisers that experience widely prevalent climate change-related scenarios, including deoxygenation. Generally, she is interested in climate change microbiology, microbial ecology, and the ecophysiology of microorganisms.
About this blog section
The section #FascinatingMicrobes for the #FEMSmicroBlog explains the science behind a paper and highlights the significance and broader context of a recent finding. One of the main goals is to share the fascinating spectrum of microbes across all fields of microbiology.
| Do you want to be a guest contributor? |
| The #FEMSmicroBlog welcomes external bloggers, writers and SciComm enthusiasts. Get in touch if you want to share your idea for a blog entry with us! |



