To fully understand the impact of microbes on our lives, we need to better understand the functions of their proteins. With the explosion of omics data, we now have the basics to characterize the vast amounts of prokaryotic proteins. The review “Elucidating the functional roles of prokaryotic proteins using big data and artificial intelligence” in FEMS Microbiology Reviews provides a primer for microbiologists on AI-based methods. Anne-Kristin Kaster and Sagarika Chakraborty explain for the #FEMSmicroBlog how data-based approaches can help make sense of the microbial world. #FascinatingMicrobes
Traditional data annotation methods are not enough
Bacteria and Archaea play critical roles in global biogeochemical cycles and have numerous applications in biotechnology. Annotating protein sequences according to their biological functions is one of the key steps in understanding microbial diversity, metabolic potential and evolutionary history.
Experimental methods to identify protein functions are however time-consuming and expensive, leaving many proteins uncharacterized even today. This means that even the best studied microorganisms are still not fully functionally characterized.
One example is the model organism Escherichia coli — the “favorite pet” of microbiologists for over 100 years. Only about 70% of its proteins are functionally characterized so far while more than 2% of protein-encoding genes are not characterized at all.
These uncharacterized, so-called hypothetical, proteins are referred to as “functional dark matter”. They are found across all microbial species and represents an enormous knowledge gap; a problem that calls for more automated and scalable approaches.
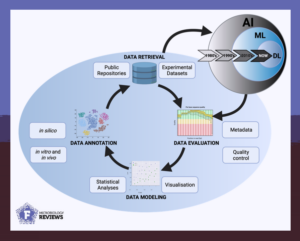
The next step: AI for functional characterization of proteins
In recent years, the integration of big data and artificial intelligence has emerged as a promising approach to predict and facilitate the discovery of protein functions. The review “Elucidating the functional roles of prokaryotic proteins using big data and artificial intelligence” in FEMS Microbiology Reviews delves into different types of machine and deep learning techniques, which have been developed for protein annotation.
To recognize patterns within large datasets, trained models use statistical algorithms. These can incorporate diverse data sources such as sequence homology, structural information and functional annotations from related proteins.
As more genomic data become available, machine learning and deep learning models will play increasingly important roles in identifying novel genes and characterizing the functions of individual proteins within microorganisms from different habitats. Data-based approaches can overcome current limitations and hopefully open the door to an unexplored world in microbial genomes.
Nevertheless, the direct experimental analysis will still remain the gold standard of functional protein annotation for the foreseeable future. An experimental feedback loop is therefore of great importance to further test and train the models for accurate predictions.
The continued collaboration between computational biologists and experimentalists will be necessary to validate and refine approaches and to ensure that they are applicable to a wide range of microbial systems.
- Read the article “Elucidating the functional roles of prokaryotic proteins using big data and artificial intelligence” by Ardern et al. (2023).
About the authors of this blog
Anne-Kristin Kaster is the principal investigator for Microbial Genetics and Bioinformatics at the Institute for Biological Interfaces In Karlsruhe.
Sagarika Chakraborty is a PhD student in Anne-Kristin Kaster’s research lab.
About this blog section
The section #FascinatingMicrobes for the #FEMSmicroBlog explains the science behind a paper and highlights the significance and broader context of a recent finding. One of the main goals is to share the fascinating spectrum of microbes across all fields of microbiology.
| Do you want to be a guest contributor? |
| The #FEMSmicroBlog welcomes external bloggers, writers and SciComm enthusiasts. Get in touch if you want to share your idea for a blog entry with us! |



