Our academy journal microLife has announced the winners of its Article Awards for the best article published in 2022! Our congratulations go to Anamaria Babosan, Louna Fruchard, Evelyne Krin, André Carvalho, Didier Mazel and Zeynep Baharoglu who wrote the excellent research article “Nonessential tRNA and rRNA modifications impact the bacterial response to sub-MIC antibiotic stress”.
All authors who published in microLife in 2022 were automatically considered for this award upon submission of their paper. The best article was awarded with €1,000 which was split equally between all the authors of the winning paper.
We interviewed all the authors to find out more about the inspiration behind this article:

Could you provide a brief, simple overview of the topic your paper covers?
The aim of this study was to uncover new genes involved in survival and growth upon exposure of bacteria to low doses of antibiotics, below sub-minimal inhibitory concentrations (sub-MICs), in Vibrio cholerae.
Using a high-throughput method, the impact of disruption of each gene on the ability of bacteria to thrive in the presence of non-lethal doses of various antibiotics was determined.
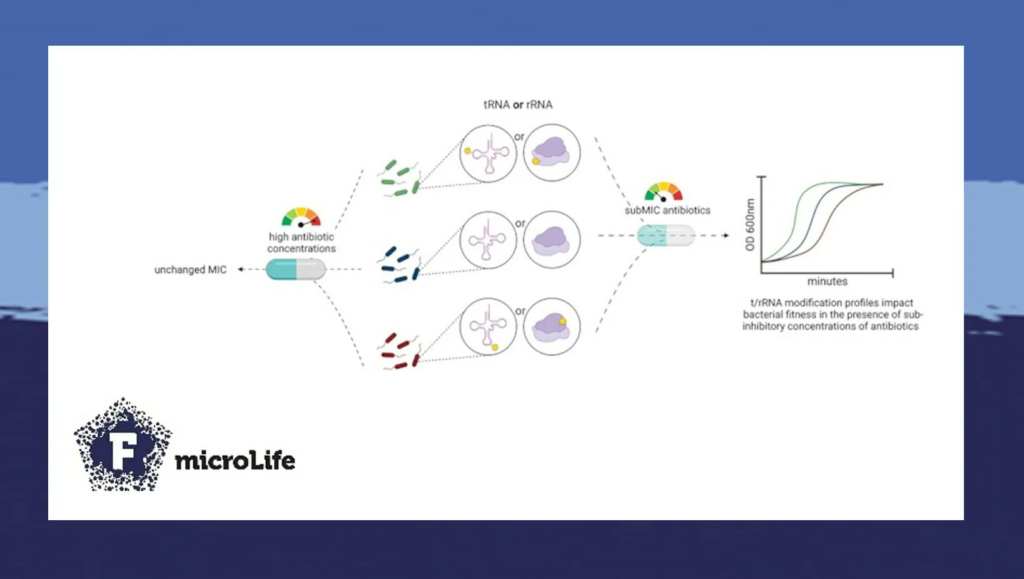
The results reveal a central role of transfer RNA (tRNA) and ribosomal RNA (rRNA) modifications in the response to antibiotic stress. These non-coding RNAs are the main actors of messenger RNA translation into proteins: rRNAs being important constituents of ribosomes and tRNA delivering the relevant amino-acid to the protein being synthesized. t/rRNA modifications influence translation efficiency and fidelity, and therefore, the bacterial proteome composition. While some tRNA modifications are essential for viability, the absence of most shows no phenotype in unstressed cells.
Our study identified RNA modification genes whose inactivation shows no fitness defect in standard growth conditions, but with strong fitness defect or advantage in response to antibiotic stress. Moreover, the identified genes had not been previously associated with antibiotic resistance. We found that their inactivation confers increased or decreased tolerance to antibiotics, allowing further study of involved factors. The fact that different RNA modifications have opposite effects on tolerance to different antibiotics highlights the complexity of such a network.
RNA modification profiles may thus be altered in bacteria by antibiotic pressure and that resulting variations in protein synthesis could be critical factors involved in the bacterial response to stress.
For most of the tested t/rRNA modification genes mutants, survival profiles after a 20h-long lethal antibiotic treatment correlate with their fitness profiles in sub-MICs of antibiotics, while their resistance is globally unchanged compared to the wild-type strain, which is a hallmark of antibiotic tolerance.
The results from our study suggest an epitranscriptomic response to antibiotics, and will help further understand bacterial pathways mobilized in the response to antibiotics and new functions essential for survival.”
What do the results reveal about possible directions of antibiotic development?
Thorough characterization of the effects of each RNA modification on the expression of specific stress response factors may lead to the development of antibiotics against these new targets, or to strategies inhibiting the translation of specific proteins.
Additionally, the effects of several RNA modifications on antibiotic tolerance (the rate in which they are killed by antibiotics), without necessarily affecting resistance (MIC), can lead to their study as markers of relapsing infections. Antibiotic tolerance and persistence are potentially significant contributors to infection relapse.”
What encouraged you to perform research in this area of microbiology?
While antibiotic resistance is now an important public health issue, it has become clear that exposure of bacteria to lower, non-lethal doses of antibiotics is a source of stress which prompts bacteria to adopt survival strategies. These survival strategies can be in the origin of resistance development, and we aim to thoroughly characterize the stress responses of bacteria to antibiotic concentrations below the minimal inhibitory concentrations (sub-MIC). This approach studying factors important for growth in presence of sub-MIC antibiotics also allowed us to uncover factors responsible of tolerance to antibiotics. Among those were RNA modification factors. t/rRNA modifications and their physiological effects have been mostly studied in eukaryotes and linked to human diseases. In bacteria, literature about the involvement of RNA (especially tRNA) modifications, in stress response and adaptation to antibiotics, is less abundant.”
What do you see as the next steps in this area of research?
Since RNA modifications affect protein synthesis, it is important to address how differential translation associated with RNA modifications can generate diversity in the abundance of specific proteins, and eventually adaptation to various environments. The diversity of RNA modifications, their specific effects on various proteins and cellular processes and the possibility that their expression may be regulated by various stresses make them now a promising field of study.
With the development of RNA modification detection methods such as NGS-based techniques and especially Nanopore sequencing, we believe this field will be a hot topic for the next years.
One can also envisage applying identified associations between specific RNA modifications and antibiotic resistance phenotypes, for the analysis of available bacterial collections with established genome sequences and AMR profiles, for which observed phenotypic resistance does not always correlate with known resistance factors.”
Read the 2022 award winning paper: Nonessential tRNA and rRNA modifications impact the bacterial response to sub-MIC antibiotic stress



