S. Soldatou, G. H. Eldjarn, A. Huerta-Uribe, S. Rogers and K. R. Duncan: Winners (2019) of the MiniReview Award from FEMS Microbiology Letters
27-01-20
Sylvia Soldatou, Grimur Hjorleifsson Eldjarn, Alejandro Huerta-Uribe, Simon Rogers and Katherine Duncan are the winners of the 2019 MiniReview award from FEMS Microbiology Letters. Their winning article is: Linking biosynthetic and chemical space to accelerate microbial secondary metabolite discovery.
You can follow the authors on Twitter: Sylvia Soldatou: @SylviaSoldatou; Grimur Hjorleifsson Eldjarn: @Grimur_H; Alejandro Huerta-Uribe: @alex_hu_; Simon Rogers: @sdrogers; Katherine R. Duncan: @kate_duncan

The paper recieved the most votes from the editorial board and is fully Open Access. We interviewed the authors to find out more about the inspiration behind their paper and they took the time to consolidate their answers:
Could you provide a brief, simple overview of the topic your paper covers and why is it important?
Microbial genome sequencing has uncovered the incredible potential of strains to produce medically-relevant chemistry (often termed secondary or specialised metabolites). This biosynthetic capacity has been underpinned by the in-silico prediction of sequence regions (known as Biosynthetic Gene Clusters; BGCs) that encode the enzymes for these potential metabolic products. Researchers can therefore mine genome sequences for putative BGCs whilst simultaneously growing the strains and analysing their metabolites. Despite computational tools existing for individual datasets, the linking of these rich and complex biosynthetic and chemical data is largely a manual process. This is fundamental bottleneck for the discovery of new chemistry and one that is essential to overcome to enable new biosynthetic and chemical discovery against the increasing global threat of strains resistant to currently available antibiotics.
In our review paper we initially describe the creation of datasets for the prediction of BGCs and metabolites. This is followed by approaches to link these metabolomic and genomic datasets. Firstly, our review focuses on targeted (chemical investigation as a result of a particular BGC of interest) and multi-targeted (multiple reasons for chemical investigation in addition to genome mining) approaches, which are more common.
Our review then defines automated linking approaches and we split this into feature-based (prediction of structural properties), correlation-based (matching patterns of source strain occurence of genes and chemistry) and hybrid approaches. All methods are evidenced with multiple examples, showing the utility and application of the approaches discussed. We finish the review discussing methods to validate links between genes and chemistry and also the current challenges and future direction of this exciting research area.”
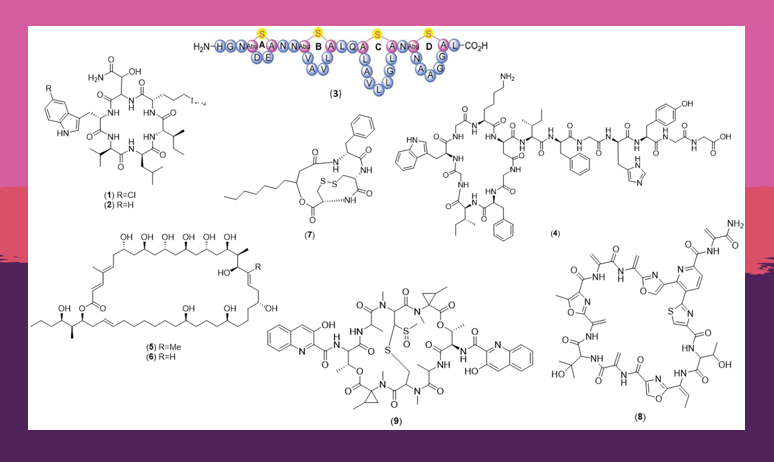
What encouraged you to review research this area of microbiology?
Actinobacteria, a phyla of bacteria, are unsurpassed in the production of bioactive microbial metabolites, many of which have been developed to drug candidates such as chloramphenicol and streptomycin produced by Streptomyces spp.
Streptomyces represents the largest genus of Actinobacteria and has shown enormous potential for the field of drug discovery due to the remarkably large genome and the ability to produce structurally diverse metabolites with promising biological activities. However, natural products drug discovery is easier said than done; traditional isolation techniques such as analytical chemistry methodologies can be time consuming, there is a high rediscovery rate and it is difficult to predict which BGCs will be expressed under laboratory conditions.
The golden age of antibiotics has long passed and a new approach which combines the biosynthetic capabilities of microorganisms (genome mining) with comparative metabolomics is needed. With recent developments in computational methods to recognise features and similarities in both mass spectrometry data and gene clusters, microbial drug discovery is once again both exciting and promising. Computational approaches are illuminating previously unknown areas of the microbial chemical space and have the potential to greatly accelerate discovery from many microorganisms, not only Actinobacteria.”
What do you see as the next steps in this area of research?
Further improvements in instrumentation make larger and more complete data sets, covering more strains and modalities, possible.
We believe that larger and larger datasets will be produced in the coming years. These, in turn, will make improvements in data processing algorithms necessary. In particular, we believe that these datasets will be amenable to machine learning approaches, further establishing this computational-era of natural product-based drug discovery.
Collaborative efforts are also required in order to increase the coverage and quality of the available data sets, as well as their interpretation. This includes both interdisciplinary collaboration between chemists, microbiologists and computational scientists, but also increased data sharing and development of open access platforms and databases for the scientific community.”
Read the 2019 award winning MiniReview paper: Linking biosynthetic and chemical space to accelerate microbial secondary metabolite discovery
See more FEMS Journals Article Awards



