Faecal pollution of water resources can pose a serious health risk to water consumers due to the possible presence of enteric pathogens. Genetic methods, such as qPCR and sequencing, have revolutionised the way we detect and analyse microbial faecal pollution in water, as outlined in the review “Have genetic targets for faecal pollution diagnostics and source tracking revolutionised water quality analysis yet?” in FEMS Microbiology Reviews. Katalin Demeter and Andreas Farnleitner explain for the #FEMSmicroBlog how genetic methods can support health-related water microbiology research. #MicrobiologyIsEverywhere
Detecting faecal pollution in water samples relies on hundred-year-old methods
Despite considerable progress in the past decades, 842 000 people still die each year from diarrheal disease as a result of unsafe drinking water, sanitation, and hygiene. Adequate water, sanitation, and hygiene (WASH) management reduces health risks. Yet, there is a constant and urgent need for more comprehensive, more informative, and quicker microbiological analyses.
For well over one century, faecal pollution assessment of water has relied on the cultivation-based detection of gut commensals, such as Escherichia coli and intestinal enterococci. These faecal indicators revolutionised water quality testing and public health protection at the end of the 19th century.
Despite great technological advancements in the life sciences, water, sanitation, and hygiene (WASH) management still relies heavily on the biological detection of faecal indicators.
Genetic methods offer unprecedented opportunities
Recent advances in DNA sequencing revealed the immense richness and diversity of the microbial world. We now know that gut microbiotas profoundly differ from free-living microbial communities across the biosphere, and that gut microbiotas of animal species differ immensely, with both host phylogeny and diet being key drivers.
Novel molecular biological and genetic tools offer promising ways to analyse and track faecal microorganisms in water. PCR and qPCR assays have been developed for various general and host-associated bacterial and viral DNA targets. High-throughput DNA sequencing allows analysing of the faecal signature of the aquatic water microbiome.
Genetic faecal pollution diagnostics: a revolution
The systematic review “Have genetic targets for faecal pollution diagnostics and source tracking revolutionised water quality analysis yet?” in FEMS Microbiology Reviews outlines the application areas, advantages, and limitations of genetic methods for faecal pollution analysis in water. The review relies on more than 1,100 publications, including 649 application studies.
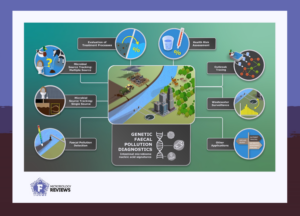
The analysis revealed seven main areas in which genetic methods are applied to analyse faecal pollution of water samples. These are Evaluation of Treatment Processes, Health Risk Assessment, Outbreak Tracing, Wastewater Surveillance, Microbial Source Tracking from single or multiple sources, and Faecal Pollution Detection.
Given the consistency of methods and assessment types, the review defines this emerging scientific field as a new discipline and coins it as Genetic Faecal Pollution Diagnostics (GFPD). GFPD has already revolutionised faecal pollution detection and microbial source tracking through general and host-associated faecal markers. As such, this new approach gets results faster while it is also able to identify the pollution source. Notably, three areas, faecal pollution detection and microbial source tracking (from single or multiple sources), encompass almost 70% of identified application studies.
Genetic methods: a powerful but not magical toolkit
Results from genetic methods provide more information on the faecal pollution characteristics of water samples than cultivation-based methods do. Additionally, storing DNA extracts allows compiling of large sample sets into databases and easily transporting samples for collaborations.
However, detecting genetic material does not provide sufficient information on the viability or infectivity of the identified organisms. This information is very important in some application areas, such as disinfection efficiency assessment. Here, cultivation-based methods may still be a crucial tool.
The comprehensive meta-analysis presented in “Have genetic targets for faecal pollution diagnostics and source tracking revolutionised water quality analysis yet?” in FEMS Microbiology Reviews provides the scientific status quo of faecal pollution analysis of water resources. It also discusses the benefits and challenges of genetic analysis methods in GFPD and serves as comprehensive source of information for microbiologists, water hygienists, and water management professionals.
- Read the review “Have genetic targets for faecal pollution diagnostics and source tracking revolutionised water quality analysis yet?” in FEMS Microbiology Reviews by Demeter et al. (2023).
- Read: #FEMSmicroBlog: Wastewater samples to predict viral outbreaks
- Read: #FEMSmicroBlog: From drain to data – tackling local COVID-19 outbreaks with wastewater testing
- Read: #FEMSmicroBlog: A simplified, RNA-based workflow for SARS-CoV-2 detection in wastewater
About the authors of this blog
 Katalin Demeter is a postdoctoral researcher and Andreas H. Farnleitner full professor and leader of the Research Group of Microbiology and Molecular Diagnostics at the TU Wien and head of the Division Water & Health at the Karl Landsteiner University Krems. Katalin’s primary research interest is microbial source tracking, while Andreas’ field encompasses diverse questions of health-related water microbiology. Both are members of the Austrian Interuniversity Cooperation Centre Water & Health. Here, they are pictured among the co-authors of the systematic review (second and third from right in the bottom row).
Katalin Demeter is a postdoctoral researcher and Andreas H. Farnleitner full professor and leader of the Research Group of Microbiology and Molecular Diagnostics at the TU Wien and head of the Division Water & Health at the Karl Landsteiner University Krems. Katalin’s primary research interest is microbial source tracking, while Andreas’ field encompasses diverse questions of health-related water microbiology. Both are members of the Austrian Interuniversity Cooperation Centre Water & Health. Here, they are pictured among the co-authors of the systematic review (second and third from right in the bottom row).
About this blog section
The section #MicrobiologyIsEverywhere highlights the global relevance of microbiology. The section acknowledges that microbiology knows no borders, as well as the fact that microbiologists are everywhere and our FEMS network extends well beyond Europe. This blog entry type accepts contributions from excellent blogs translated into English. Regional stories with global relevance are welcomed. National or international events sponsored, organised or connected to FEMS are also covered.
| Do you want to be a guest contributor? |
| The #FEMSmicroBlog welcomes external bloggers, writers and SciComm enthusiasts. Get in touch if you want to share your idea for a blog entry with us! |



